Welcome to the TassDB2
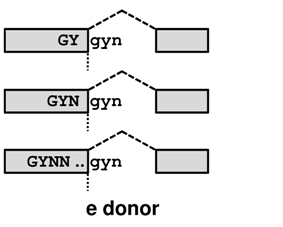
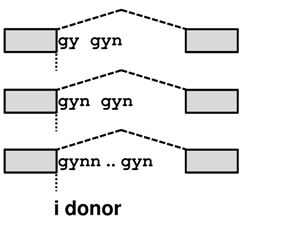
TassDB2 contains data about GY(N)0-10GYN
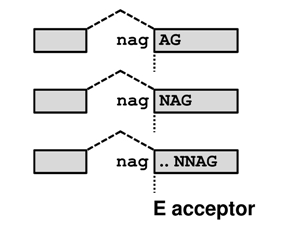
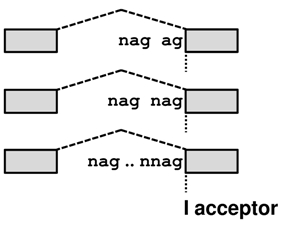
donors and NAG(N)0-10AG acceptors in human and mouse, where the distance between the tandem splice sites ranges from 2 to 12. It stores information about 1,329,181 splice sites, 14,312 of which are supported by EST/mRNA evidence.
Nomenclature for the splice sites and transcripts
Tandem alternative splicing using the intron-proximal site (E acceptor / e donor - the distal part of the tandem becomes EXONIC) results in the E/e transcript, whereas splicing using the intron-distal site (I acceptor / i donor - as the entire tandem becomes INTRONIC) in the I/i transcript. For an illustration of these effects move your mouse over the transcript names in the picture below.
TassDB allows to:
- search for all tandem splice sites of a human or mouse gene (or group of genes) given its symbol (HGNC or MGI conventions), an alias name, and/or transcript accession number
⇒ Quick search
- search for human or mouse genes containing tandem splice sites with specific features, e.g.
- distance between a pair of tandem sites
- number of ESTs/mRNAs that match both splice forms
- conservation in four species
- splice site scores
- location in the UTR or in the CDS
⇒ Advanced Search